Zindaiji3 can read HDF5 file. Note that HDF5 is rather a container than a format. Zindaiji3 can handle data formatted as below.
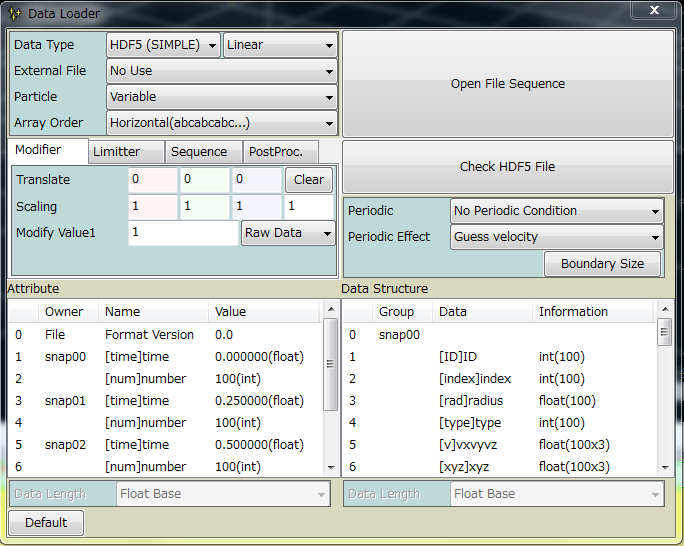


| xyz | necessary position (float) |
| vxvyvz | velocity (float) |
| axayaz | 1/2 acceleration (float) |
| jxjyjz | 1/6 jerk (float) |
| radius | radi (float) |
| dradius | derivative value of radi |
| value1 | VALUE1 (aritrary physical value) (float) |
| dvalue1 | derivative value of VALUE1 |
| type | type(int) |
| ID | ID(int) |
| index | index (int, necessary for TOKI compression) |
 Tags connected to datasets can be changed.
Tags connected to datasets can be changed.